Translate this page into:
A pilot study for diagnosis of genital Chlamydia trachomatis infections by polymerase chain reaction among symptomatic Indian women
2 Regional STD Teaching, Training and Research Center, VMMC and SJH, New Delhi, India
3 Department of Ocular Microbiology, Dr. R.P. Centre for Ophthalmic Sciences, AIIMS, New Delhi, India
4 Department of Dermatology and Venereology, AIIMS, New Delhi, India
5 Department of Biostatistics, AIIMS, New Delhi, India
Correspondence Address:
Seema Sood
Department of Microbiology, All India Institute of Medical Sciences, Ansari Nagar, New Delhi
India
| How to cite this article: Sood S, Mukherjee A, Bala M, Satpathy G, Mahajan N, Sharma A, Kapil A, Sharma VK, Pandey RM, Samantaray JC. A pilot study for diagnosis of genital Chlamydia trachomatis infections by polymerase chain reaction among symptomatic Indian women. Indian J Dermatol Venereol Leprol 2012;78:443-447 |
Abstract
Background: Chlamydia trachomatis is the most common bacterial etiology of sexually transmitted infection. Aim : A pilot study was designed using PCR for amplification and detection of a specific 517 bp sequence of the common endogenous plasmid of C. trachomatis from clinical swab specimens obtained from symptomatic female patients attending STD clinics of AIIMS and Regional STD Teaching, Training & Research Center, Safdarjang Hospital, New Delhi. Methods: 97 patients were recruited in the study, and endocervical swabs were collected following standard procedures. The samples were analyzed by PCR and direct fluorescence antibody (DFA) for detection of C. trachomatis, and the sensitivity, specificity, PPV and NPV of PCR were calculated taking DFA as gold standard. Results: Out of 97 samples tested, 9 were positive for C. trachomatis by PCR. 1 PCR positive patient was negative by DFA although a total of 11 patients were positive by DFA. The sensitivity, specificity, PPV and NPV of PCR with reference to DFA was 72.73%, 98.84%, 88.89% and 96.59%, respectively. This PCR had high specificity and NPV for detection of C.trachomatis. Conclusions : In light of the introduction of enhanced syndromic approach, which involves the use of laboratory techniques (wherever possible) to confirm clinical diagnosis, a diagnostic PCR with high specificity and NPV is particularly valuable for determination of etiological diagnosis and hence contribute to judicious use of antimicrobials in the community.Introduction
Chlamydia trachomatis is the commonest bacterial sexually transmitted disease worldwide. [1] WHO estimates that about 92 million cases occur annually on a global basis and 43 million in South East Asia alone, attesting to its public health importance. [2] The diagnosis often poses serious problems as approximately 70% - 80% of women and up to 50% of men are asymptomatic. [3],[4] Undiagnosed infections often present as pelvic inflammatory disease, leading to ectopic pregnancy, infertility or other adverse health outcomes in women, [5] inclusion conjunctivitis and interstitial pneumonia in newborn. [6],[7],[8] In addition, Chlamydia infection appears to increase the risk of HIV transmission 2 to 4 times during sexual intercourse. [9],[10] Further, it has also been implicated as a risk factor for developing cervical carcinoma. [2] Therefore, an urgent need is felt to actively diagnose the disease because while on one hand, it can easily be treated by oral antibiotics, untreated cases will lead to an increased duration of infectivity and transmissibility and long term sequelae, especially in women.
WHO recommends the syndromic approach as a low cost feasible alternative to the laboratory testing for STIs. The use of this approach has shown to be effective in the treatment of genital ulcer syndrome, and works well in other situations like the management of opthalmia neonatorum and symptomatic men. However, the use of this approach in the management of female patients with vaginal discharge is contentious. [9],[10]
Reliable estimate of prevalence data for genital Chlamydia infection using sensitive and specific techniques like nucleic acid amplification tests is lacking in India. [11] However, the few available reports describe an increased incidence of genital Chlamydial infection, especially in Indian women. [12],[13] Therefore, the present pilot study was designed to standardize an in-house PCR and to determine the validity statistics of PCR with respect to DFA. The gene targeted for in the present study was the 517 bp common endogenous plasmid. The advantage of the plasmid gene as PCR target is its presence in high numbers (7 - 10/organism) and hence an increased sensitivity as compared to PCR targeting the Major Outer Membrane Protein (MOMP) gene, the other commonly used target. Further, the chosen sequence was specifically checked for absence of the 377 bp region whose deletion has been reported from Sweden and has led to the inability of the commercially available kits to detect these strains, called the new variant Chlamydia trachomatis (nvCT). However, the in-house PCR used by us would detect all cases, including those with the above-mentioned deletion.
Methods
Sample size calculation
The sample size was calculated based on the results of previous published data in which the sensitivity of PCR was found to be about 75%.
Taking the sensitivity of the gold standard (DFA) to be 90%, a sample size of 94 had 80% power and a 2-sided confidence limit of 95% to detect this difference in sensitivity of about 15%.
Hence, a total of 97 symptomatic female patients, presenting to the STD clinic and Dermatology OPD of AIIMS and Regional STD Teaching, Training and Research Centre, Safdarjang Hospital from January 2009 to March 2009 were included in the study. The male patients (N = 3) who presented during the same period and were also symptomatic, were not included in the study.
The inclusion criterion for these patients was presence of cervical /vaginal discharge. Menstruating women were excluded from the study.
2 endocervical swabs were collected from the above-mentioned patients. Adequate counseling of the patient and partner (if present) was done before collection .
An ethical clearance was obtained from the Ethics Sub- Committee, All India Institute of Medical Sciences, New Delhi.
Microbiological investigations
DFA: Using the first swab, a bedside smear was made on a clean glass slide and was processed subsequently as per manufacturer′s instructions (Syva Microtrak, California, USA) for Direct Fluorescent Antibody (DFA). Slides were read using a fluorescent microscope (Nikon)-a 40X objective was used for screening, and a 100X objective was used for confirmation of morphology. Slides were examined for apple-green colored elementary bodies contrasted against the reddish-brown background of the counterstained cells. The presence of > 10 such structures in a slide were taken to be positive. If elementary bodies were <10 in number per slide, it was taken as doubtful positive while when no such structures were seen, the sample was considered negative.
PCR: The second swab was stored at -20° C for PCR targeting the common endogenous plasmid. DNA was extracted using the QIA amp DNA mini kit (from Qiagen) according to manufacturer′s instructions.
Bacterial isolates and control: A panel of strains comprising of N. gonorrhoeae (WHO-C strain), E. coli (ATCC 25922), S. aureus (ATCC 25923), Pseudomonas aeruginosa. (ATCC 27853), N. sicca (ATCC 29193) and Klebsiella pneumoniae (ATCC 700603) were taken for specificity studies. Positive control for Chlamydia trachomatis was DNA from a known positive patient sample.
For PCR, primers designed by Class et al., (1991) targeting the common endogenous plasmid, were used. The forward primer 5′GGA CAA ATC GTA TCT CGG′3 and reverse primer 5′ GAA ACC AAC TCT ACG CTG 3′ was used. Further, an inhibition in the samples was checked by PCR for β-globin gene. This gene serves as an internal control for PCR reaction as β-globin gene is constitutively expressed in all human cells and hence its presence indicates presence of human DNA (thus checking for an adequacy of extraction).
Statistical analysis
In our study, PCR was compared with DFA as gold standard. Similar approach has been adopted by other studies as well. [3] Statistical analysis were done using the SPSS 17 software.
Results
The age of the 97 patients ranged from 15 to 50 years with an average of 29.7 years (S.D.=6.38); 2 could not specify their age. The study sample was an essentially younger one, with 73% patients between 15 - 34 years of age. The presenting complaint of 89 (91.75%) of the patients was vaginal discharge alone. Of the remaining female patients, 1 had complaints of primary infertility, 3 had complaints of secondary infertility, 3 had vaginal discharge with IUCD, and 1 patient had an intense pruritus with herpes in addition to vaginal discharge.
Out of the 97 smears tested by DFA, 11 samples were positive for Chlamydia trachomatis [Figure - 1] while 9 were doubtful positive.
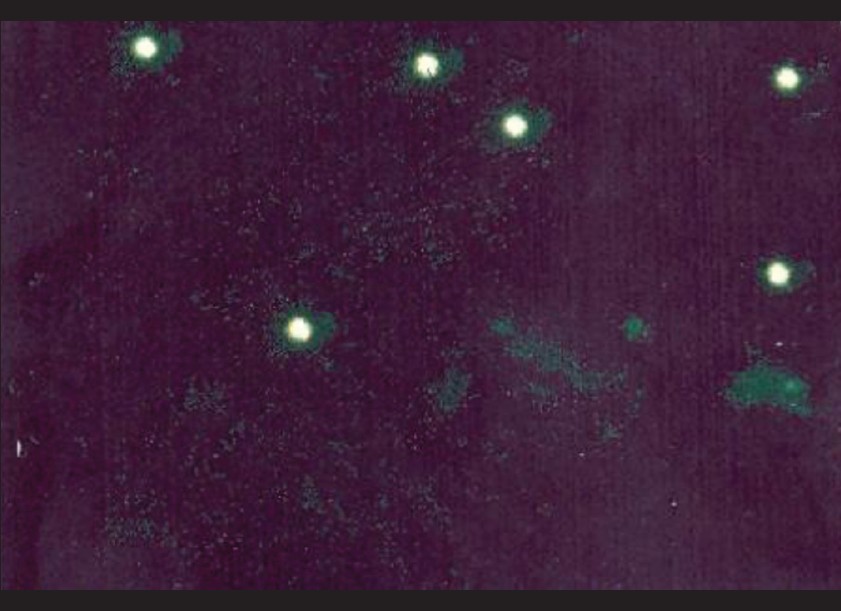 |
| Figure 1: Apple-green EB bodies against the reddish-brown background of the counterstained cells in a sample positive for Chlamydia trachomatis by DFA (100X) |
PCR for Chlamydia trachomatis: Out of the 97 samples tested , 9 were positive for C. trachomatis by PCR. [Figure - 2] shows the gel picture of a positive clinical sample for C.trachomatis along with positive control.
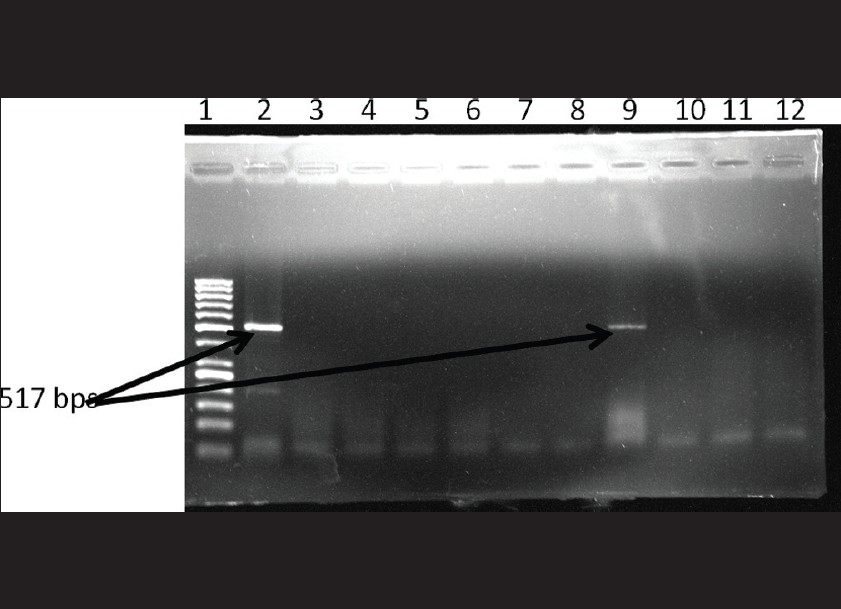 |
| Figure 2: Gel picture showing positive clinical sample along with positive control. Lane 1: 50 bp DNA ladder Lane 2: positive control Lane 9: Sample positive for Chlamydia trachomatis Lane3-8, 10-12: Sample negative for Chlamydia trachomatis |
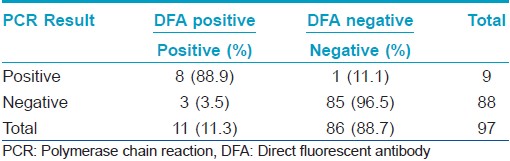
Association of DFA and PCR results: PCR for C. trachomatis was positive in 8 out of the 11 DFA positive patients (72.73%). 1 PCR positive patient was negative for DFA. No doubtful positive sample by DFA (< 10 EB) was positive with PCR.
Validity Statistics for PCR with reference to DFA: Sensitivity, specificity, positive predictive value (PPV) and negative predictive value (NPV) of PCR with reference to DFA was 72.73% (39.32% - 92.67% with 95% C.I.), 98.84% (92.79% - 99.94% with 95% C.I.), 88.89% (50.67% -99.41% with 95% C.I.), 96.59% (89.66% - 99.12% with 95% C.I.), respectively. The percentage of false negative was 27.27%, and the percentage of false positive was 1.12%. The positive likelihood ratio (LR +) was 64.94 while the negative likelihood ratio (LR -) was 0.28 [Table - 1].
On comparing the level of DFA positivity (definite positive vs. doubtful positive) with PCR results, we found a significant association between the PCR results and the level of DFA positivity, i.e., PCR positive results are significantly associated with DFA definite positive ones (Fischers Exact Test 8.089, P value was 0.0014, as shown in [Table - 2]).

Discussion
The gold standard for diagnosis of Chlamydia trachomatis is by culture in cell lines like McCoy, Buffalo Green Monkey (BGM) or HeLa-229. [14],[15] However, tissue culture is too expensive, has low sensitivity, laborious and time consuming to be practiced in routine diagnostic laboratories. [16] In addition, it has a low sensitivity owing to issues related to adequate collection, transport, storage and processing of specimen. This has initiated search for alternative diagnostic strategies, which are more feasible in diagnostic laboratory set-up, especially in resource-restricted settings. Among the commonly used options, antigen detection by DFA has a sensitivity of around 90% and a higher specificity of 98% - 99%. The high specificity of DFA is due to its dependence on visualization of the distinctive morphology and staining characteristics of Chlamydia trachomatis inclusions and elementary bodies. Further, it detects the elementary bodies, implying the presence of active, symptomatic Chlamydia trachomatis infection. It is relatively rapid (about 30 min) and does not require refrigeration of specimens during transport. It is also used for the purpose of checking the adequacy of the swab specimen collected. However, the problem with this technique is the requirement of fluorescent microscope and an expertise for interpretation. On the other hand, PCR is also shown to be a sensitive and specific method of diagnosis. [17] Importantly, the in-house PCR developed and used in the present study is affordable as compared to the commercial PCR systems. Therefore, in the present pilot study, this in-house PCR was evaluated as compared to DFA as gold standard.
In the present study, 11 out of 97 (11.34%) samples were DFA positive while 9 out of 97 (9.28%) samples were doubtful DFA positive. The remaining 77 samples (79.38%) were negative for Chlamydia trachomatis by DFA. Similar studies conducted in India taking symptomatic male patients have found higher positives (36.6%) by DFA. [3] However, in the above-mentioned study, only patients with polymorphs present in their samples were included. A figure of 22.9% has been reported in symptomatic female patients, attending the Obstetrics & Gynecology OPD of a tertiary care hospital. This study also compares the performance of an in-house PCR with reference to DFA and found 96.7% of DFA positive samples to be positive by PCR also. [18] In our study, 8 out of 11 (72.73%) DFA positive samples were also positive by PCR. A total of 9 samples were positive by PCR, and 1 of them was negative by DFA. This apparent discrepancy can be explained by the lower sample size of our study as well as the difference in inclusion criteria. In situations like these, the CDC, Atlanta suggests several possible strategies like testing a second specimen with a different test / target, performing a different nucleic acid amplification test (NAAT), targeting a different nucleic acid sequence on the original specimen, repeating the original test on the original specimen, or bringing the patient back for a retest. However, our limited resources did not permit us to perform any of these confirmatory tests. None of the 9 doubtful (< 10 EB) positive samples were positive by PCR. There was statistically significant difference in PCR positivity between DFA positive and doubtful positive samples. Hence, it can be concluded that the cut-off of > 10 EB taken in the study is satisfactory.
The results of the present study are similar to those from Orissa where Dwivedi et al., found a prevalence of 7.04%, based on PCR in symptomatic female patients, attending the Gynecology and Obstetrics OPD of a tertiary care teaching hospital. [19] These rates were higher than those found in Mumbai (0.2%) by Brabin et al., [20] and in the multicentric trial conducted by ICMR (1.2% - 33%) by Chandok et al. [21] Other studies have documented higher rates (12.3% by Mania - Pramanik et al., and 29% by Garg et al. [22],[23] )
Studies worldwide have revealed a definite predisposition of women aged < 20 years for Chlamydia trachomatis infection. However, the present study did not reveal any particular age predisposition for Chlamydia trachomatis infection based on DFA and PCR. The probable reason may be because our study sample was an essentially younger one, with 73% between 15 - 34 years of age. The age of patients has not been significantly associated with Indian patients in other studies also. [18] This may be explained by the late onset of sexual activity in our population. Other factors like having a single partner, racial and genetic factor may also play a crucial role. [24]
The sensitivity, specificity, positive predictive value and the negative predictive value of PCR was 72.7% (range of 39.32% - 92.67% with 95% C.I), 98.84% (range of 92.79% - 99.94% with 95% C.I), 88.89% (range of 50.67% - 99.41% with 95% C.I) and 96.59 (range of 89.66% - 99.12% with 95% C.I), respectively. In light of the novel enhanced syndromic approach, which involves the use of laboratory techniques (wherever possible) to confirm the clinical diagnosis, a diagnostic PCR with high specificity and NPV will be of utmost help with respect to diagnosis and therapy, even among asymptomatic population. In addition, confirmation of etiological diagnosis will reduce widespread and empirical administration of broad-spectrum antibiotics.
| 1. |
Schachter J, Hook EW, Martin DH, Willis D, Fine P, Fuller D, et al. Confirming positive results of nucleic acid amplification tests for Chlamydia trachomatis: All NAATs are not created equal. J Clin Microbiol 2005;43:1372-3.
[Google Scholar]
|
| 2. |
Jalgaonkar SV, Pathak AA, Thakur YS. Enzyme immunoassay for rapid detection of Chlamydia trachomatis in urogenital infections. Indian J Sex Transm Dis 1990;1:23-6.
[Google Scholar]
|
| 3. |
Agrawal SK, Reddy BS, Bhalla P, Kaur H. Utility of Direct Fluorescent Antibody Test for detection of Chlamydia trachomatis and its detection in male patients with non gonococcal urethritis in New Delhi. Indian J Dermatol Venereol Leprol 2003;69:144-7.
[Google Scholar]
|
| 4. |
Vincelette J, Schirm J, Bogard M, Bourgault AM, Luijt DS, Bianchi A, et al. Multicenter Evaluation of the Fully Automated COBAS AMPLICOR PCR Test for Detection of Chlamydia trachomatis in Urogenital Specimens. J Clin Microbiol 1999;37:74-80.
[Google Scholar]
|
| 5. |
Geoffray R, Swain S. Decision analysis: Point-of care Chlamydia testing vs. laboratory based methods. Clin Med Res 2004;2:29-35.
[Google Scholar]
|
| 6. |
Stamm WE. Chlamydial infections. In: Fauci B, Kasper, Hauser, Longo, Jameson, Loscalzo, editors. Principles of Internal Medicine. 17 th ed. McGraw Hill; 2008. pg. 1070-78.
[Google Scholar]
|
| 7. |
Loeffelholz MJ, Lewinski CA, Silver SR, Purohit AP, Herman SA, Buonagurio DA, et al. Detection of Chlamydia trachomatis in endocervical specimens by polymerase chain reaction. J Clin Microbiol 1992;30:2847-51.
[Google Scholar]
|
| 8. |
Duyenhoven YT, Ossewaarde JM. Chlamydia trachomatis genotypes: Correlation with clinical manifestations of infection and patients' characteristics. Clin Infect Dis 1998;26:314-22.
[Google Scholar]
|
| 9. |
Hanson J. Assessment of sexually transmitted diseases as a risk factor for HIV seroconversion in New Orleans sexually transmitted disease clinic 1990-1998. Ann Epidemiol 2005;15:13-20.
[Google Scholar]
|
| 10. |
Korenromp E. Determinants of the impact of sexually transmitted infection treatment on prevention of HIV infection: A synthesis of evidence from Mwanza, Rakai and Masaka intervention trials. J Infect Dis 2005;191 Suppl 1:168- 78.
[Google Scholar]
|
| 11. |
George AJ, Thyagrajan S. Evaluation of diagnostic efficacy of PCR methods for Chlamydia trachomatis infection in genital and urine specimens of symptomatic men and women in India Jpn J Infect Dis 2003;56:88-92.
[Google Scholar]
|
| 12. |
Mittal A, Kapur K, Gupta S. Screening for genital Chlamydial infection in symptomatic women. Indian J Med Res 1993;98:119-23.
[Google Scholar]
|
| 13. |
Gaonkar T. Chlamydia trachomatis in women with high risk behavior: Diagnostic efficacy of non-culture tests. Indian J Sex Transm Dis 2003;24:86-90.
[Google Scholar]
|
| 14. |
Pearlman MD, McNeeley SG. A review of the microbiology, immunology, and clinical implications of Chlamydia trachomatis infections. Obstet Gynecol Surv 1992;47:448-61.
[Google Scholar]
|
| 15. |
Ward M, Ridgeway G. Chlamydia. In: Collins L, Balows A, Sussman M, editors. Topley & Wilson's Microbiology and Microbial Infections. 9 th ed. New York: Oxford University Press; 1998. p. 968-98.
th ed. New York: Oxford University Press; 1998. p. 968-98.'>[Google Scholar]
|
| 16. |
Stamm WE, Batteiger BE. Introduction to Chlamydia and Chlamydophilia. In: Mandell GL, Benett GE, Dolin R, editors. Principles and Practice of Infectious Diseases. 7 th ed. Churchill Livingstone Elsevier; 2010. p.2439-42.
[Google Scholar]
|
| 17. |
Altwegg M, Burger D, Lauper U, Schar G. Comparison of Gen-Probe PACE 2, Amplicor Roche, and a conventional PCR for the detection of Chlamydia trachomatis in genital specimens. Med Microbiol Lett 1994;3:181-7.
[Google Scholar]
|
| 18. |
Sachdeva P, Patel AL, Sachdev D, Ali M, Mittal A, Saluja D. Comparison of an in-house PCR assay, direct fluorescence assay and the Roche AMPLICOR Chlamydia trachomatis kit for detection of C. trachomatis. J Med Microbiol 2009; 58:867-73.
[Google Scholar]
|
| 19. |
Dwivedi B, Pramanik J, Sahu P, Kar S, Moharana T. Prevalence of genital chlamydia infection in females attending an Obstetric and Gynaecology out patient department in Orissa. Indian J Dermatol Venerol Leprol 2009;75:614-6.
[Google Scholar]
|
| 20. |
Brabin L, Gogate A, Gogate S, Karande A, Khanna R, Dollimore N. Reproductive tract infections, gynaecological morbidity and HIV seroprevalence among women in Mumbai, India. Bull World Health Organ 1998;76:277-87.
[Google Scholar]
|
| 21. |
Chandhok N, Datey S, Gaur L, Saxena N. Prevalence of Chlamydia trachomatis in women attending different clinics at tertiary care hospitals (An ICMR Task Force Study). J Obstet Gynaecol India 2000;53:463-7.
[Google Scholar]
|
| 22. |
Mania-Pramanik J, Potdar S, Kerkar S. Diagnosis of Chlamydia trachomatis infections. J Clin Lab Anal 2006;20:8-14.
[Google Scholar]
|
| 23. |
Garg S, Sharma N, Bhalla P, Sahay R, Raina U. Reproductive morbidity in an Indian urban slum: Need for health action. Sex Transm Inf 2002;78:68-9.
[Google Scholar]
|
| 24. |
Black CM. Current methods of laboratory diagnosis of chlamydia trachomatis infections. Clin Microbiol Rev 1997;10:160-84.
[Google Scholar]
|
Fulltext Views
1,666
PDF downloads
866





